Engineering
Life. Digitally.
An accurate and intelligent protein structure prediction platform. Accelerate drug discovery with next-generation AI models.
Our Pipelines
Existing Pipeline 1 & 2 from foldexa.bio (shown above this section)
Pipeline 1 - CDR Redesign (DiffAb) - 4-step - ~3h
Pipeline 2 - De Novo Scaffold (RFdiffusion) - 4-step - ~5h
From Target to Lead Candidate.
In Days, Not Years.
The first self-service platform that designs, validates, and scores therapeutic antibodies and engineered enzymes - with a closed-loop that gets smarter from your wet-lab results.
3-5 days
From PDB upload to ranked, developability-scored candidates. No library screening. No animal immunization.
$3,000
100+ candidates designed, 12-metric developability scored, Boltz-2 structure predicted, lab-ready sequences.
100%
Hit rate on anti-Tie2. 10/10 designs passed. Published in mAbs (2025). pLDDT 0.96, ddG -25.75.
DiffAb / RFdiff
RL-Guided Gen
Pre-trained weights
ProteinMPNN
Sequence Opt.
Pre-trained weights
Boltz-2 + Protenix
Structure + Affinity
Pre-trained ensemble
12-Metric Cascade
Developability
Jain et al. benchmarks
Pareto Ranking
Multi-objective
Evidence-based wts
Wet Lab -> RL
Feedback Loop
CPU model, 10s
6-step pipeline - All models use pre-trained weights (inference only) - RL feedback via lightweight CPU model
Biotech R&D Teams
The problem
Cannot afford $500K CRO campaigns. Investors want de-risked candidates before committing to wet lab.
What Foldexa gives you
100+ validated candidates for $3K. Boltz-2 affinity + 12-metric scores for investor decks.
University Labs
The problem
Need therapeutic antibodies on grant budget. Publication deadline in 6 months.
What Foldexa gives you
Publishable structure predictions in a week. DiffAb, Boltz-2, ProteinMPNN all citable.
Pharma Discovery
The problem
95% developability failure rate. Late-stage candidates fail on immunogenicity, aggregation.
What Foldexa gives you
Every candidate pre-screened against 137 clinical-stage mAbs (Jain PNAS 2017).
Enzyme Engineering
The problem
PET-degrading enzymes too slow to screen for thermostability variants.
What Foldexa gives you
Enzyme mode: Tm optimization, active site conservation, solubility. 200 variants/round.
$3,000 per target - Results in 3-5 days - No GPU infrastructure needed
From Algorithms
to the Cure.
Foldexa was created by four enthusiasts: Azamat, Kanat, Issabek and Rauan with a vision to democratize protein science.
We combine state-of-the-art diffusion models (DiffAb, RFdiffusion) with AlphaFold2 to create a seamless pipeline for de novo antibody design.
Our Pipelines
Simple. Powerful. Fast.
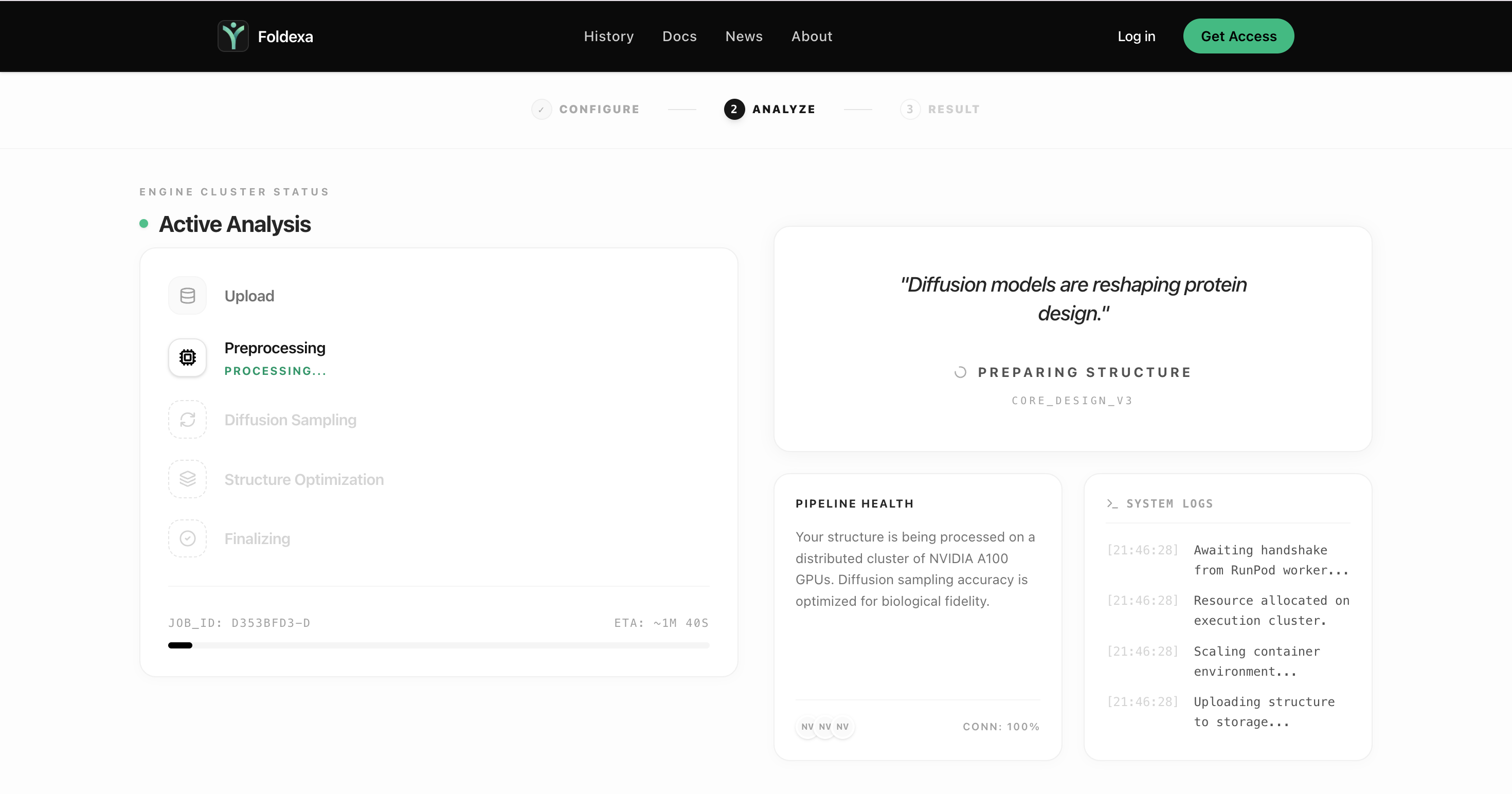
Step 03
Generate
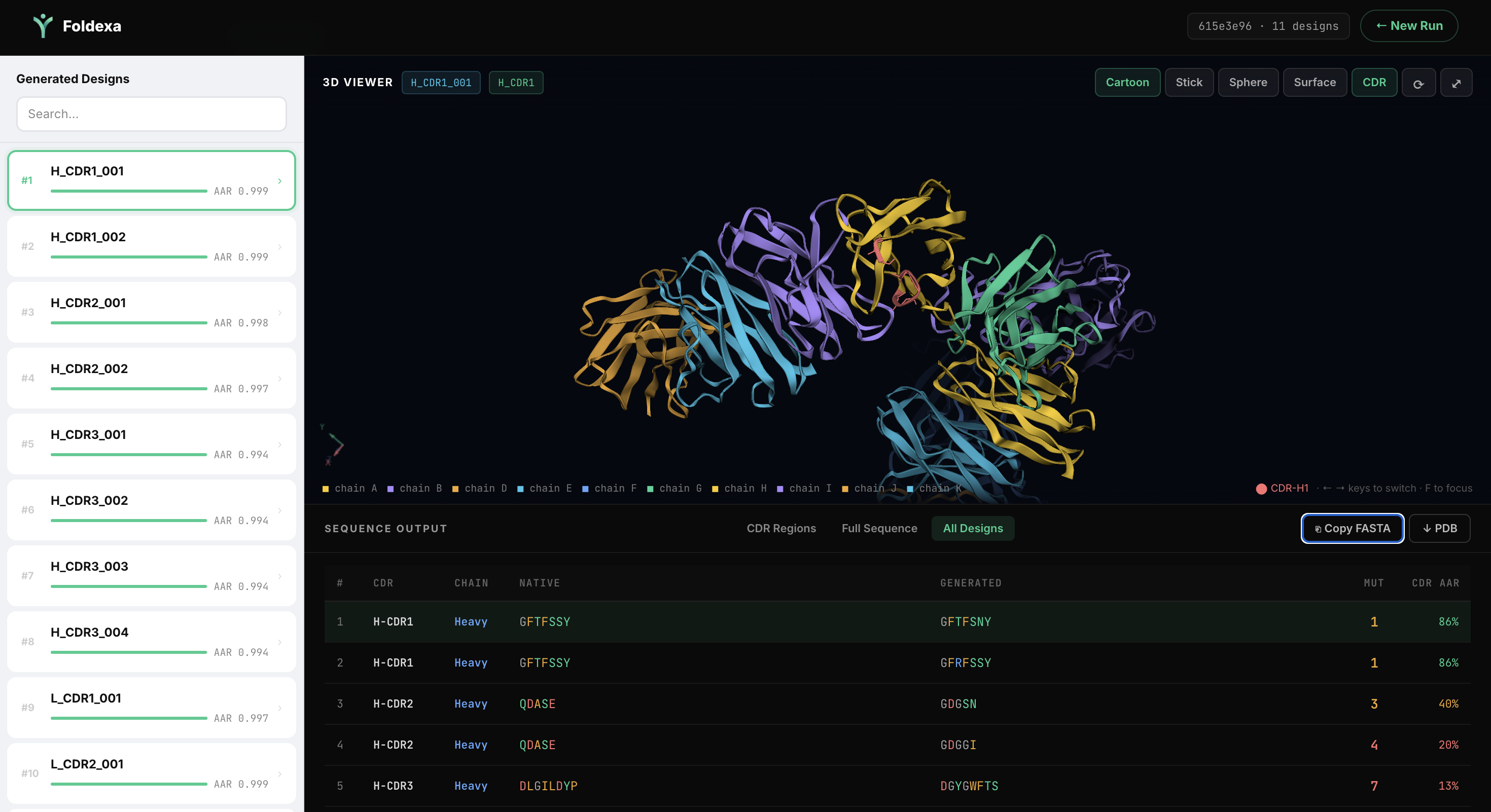
Step 04
Analyze Results
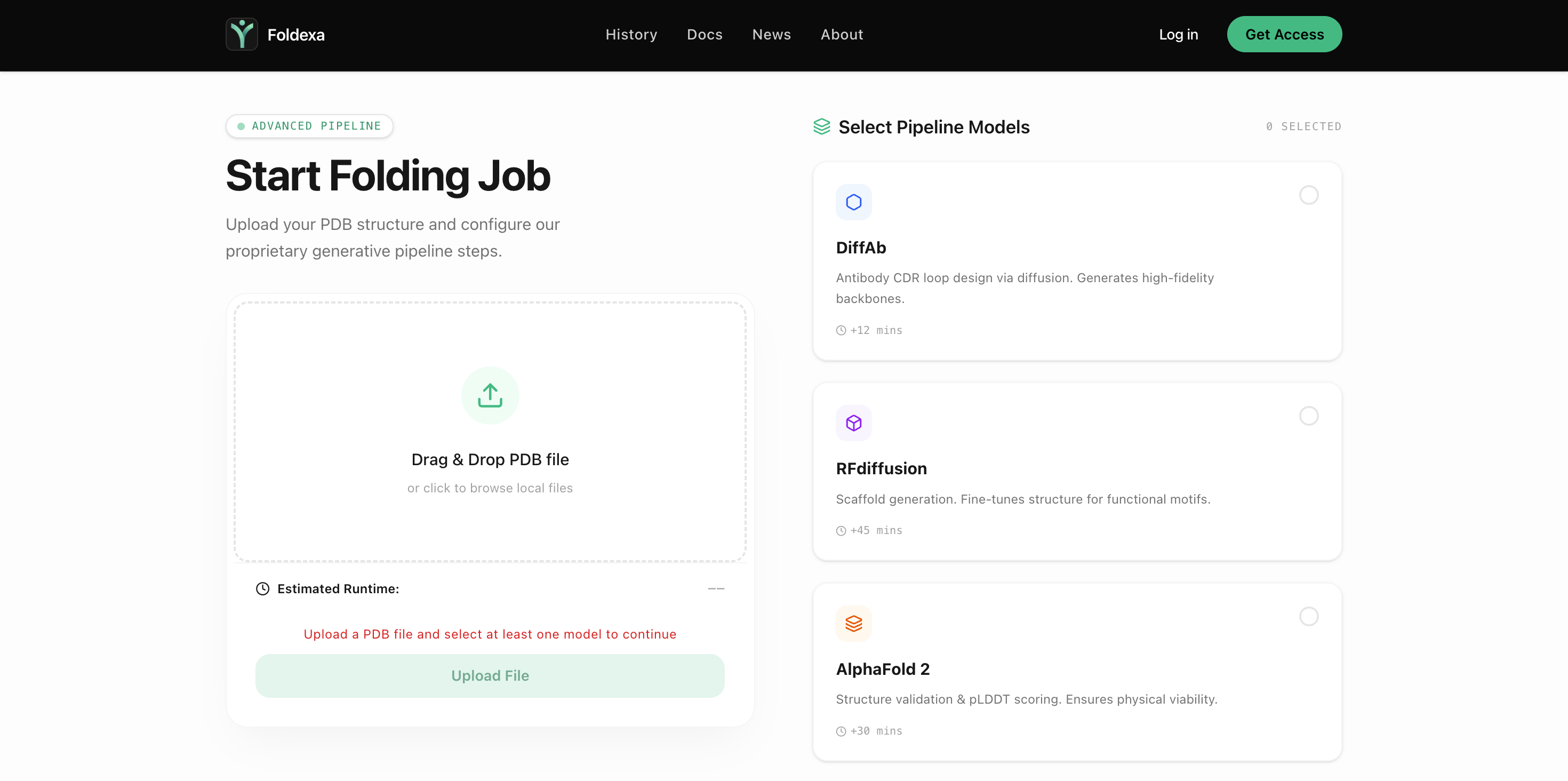
Step 01
Upload Structure
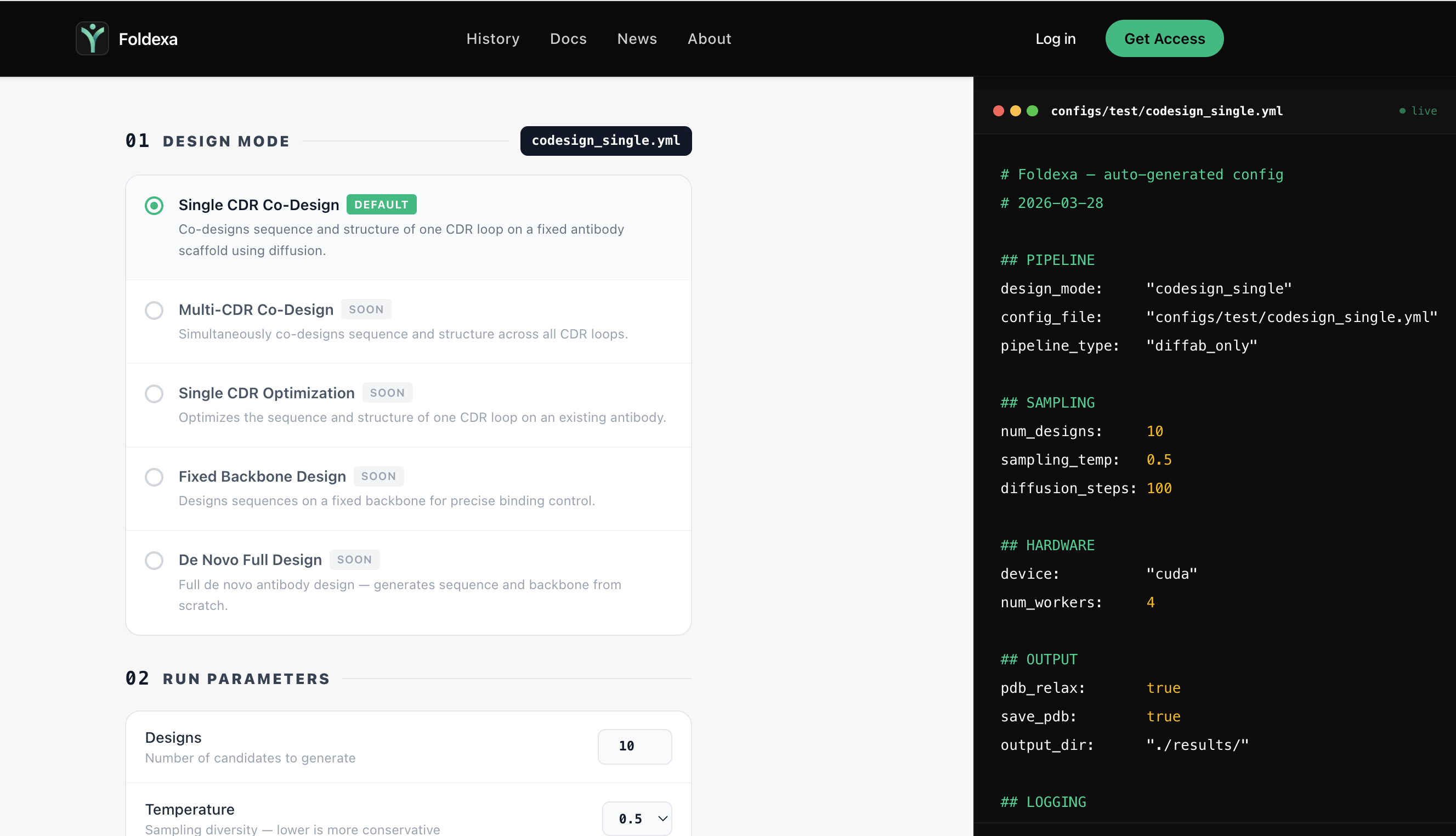
Step 02
Choose Pipeline

Step 03
Generate

Step 04
Analyze Results

Step 01
Upload Structure

Step 02
Choose Pipeline
Pipeline Benchmark vs.
World-Class Platforms
Anti-Tie2 CDR redesign · AF2-multimer predicted metrics · In Silico metrics · February 2026
Core Metrics Comparison
Interface PAE (i_PAE)
Lower = higher confidence in binding pose
Foldexa.bio
Best (#1)
Foldexa.bio
Top-10 avg
Reference
hTAAB-hTie2
Chai-2
(median)
CDR-L1 Backbone RMSD
Angstroms (Å) — lower is better
Foldexa.bio
Best (#1)
Foldexa.bio
Top-10 avg
Reference
hTAAB-hTie2
DiffAb
(scRMSD)
Binding Free Energy (Rosetta ddG)
kcal/mol — more negative = stronger binding
Foldexa.bio
Best (#4)
Foldexa.bio
Top-10 avg
RFdiffusion
(Bennett)
Reference
hTAAB-hTie2
Key Performance Indicators
(10/10 designs passed)
(near-native)
(kcal/mol)
(interface confidence)
(structure quality)
Pipeline Feature Comparison
| Feature | Foldexa.bio | RFdiffusion | DiffAb | Chai-2 |
|---|---|---|---|---|
| CDR redesign | ✓ All CDRs | ✓ | ✓ | — (structure only) |
| Hit rate | 100% (10/10) | ~30–60% | ~25% | N/A |
| Structural validation | AF2-multimer | AF2 / ESMFold | RMSD only | Built-in |
| Binding energy scoring | Rosetta ddG | Rosetta / PyRosetta | — | — |
| End-to-end pipeline | ✓ Automated | Manual assembly | Manual | Inference only |
Note: Foldexa.bio metrics from AF2-multimer predictions. Cross-platform metrics not directly comparable.
We are Foldexa
Four different paths.
One shared obsession.
We came from different worlds — bioengineering, software, business, and systems architecture. But we shared one belief: protein engineering should be accessible to everyone.
Swipe to see all

Azamat Armanuly
CEOBioengineer — KAIST
"Understanding life at molecular level"
Kanat Tilekov
CTOSoftware Developer & Data Analyst
"Building the infrastructure science deserves"

Sarzmuza Issabek
COOBusiness & Growth
"Turning scientific power into real-world impact"

Rauan Bolat
CPOProject Manager & Product Developer
"Architecting systems from concept to execution"
Four disciplines. Four founders.
One platform.
Foldexa brings biology, software, and vision together to accelerate the future of protein engineering.
To democratize protein engineering and make cutting-edge AI tools accessible to researchers worldwide.
We believe that breakthrough discoveries shouldn't be limited by computational barriers.

